-Search query
-Search result
Showing all 38 items for (author: hall & dl)

EMDB-43931: 
CryoEM structure of activated CRAF/MEK/14-3-3 complex with NST-628
Method: single particle / : Quade B, Cohen SE, Huang X

EMDB-43932: 
Activated CRAF/MEK heterotetramer from focused refinement of CRAF/MEK/14-3-3 complex
Method: single particle / : Quade B, Cohen SE, Huang X

PDB-9axa: 
CryoEM structure of activated CRAF/MEK/14-3-3 complex with NST-628
Method: single particle / : Quade B, Cohen SE, Huang X

PDB-9axc: 
Activated CRAF/MEK heterotetramer from focused refinement of CRAF/MEK/14-3-3 complex
Method: single particle / : Quade B, Cohen SE, Huang X

EMDB-40856: 
Single particle reconstruction of the human LINE-1 ORF2p without substrate (apo)
Method: single particle / : van Eeuwen T, Taylor MS, Rout MP
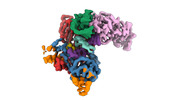
EMDB-40858: 
Structure of LINE-1 ORF2p with template:primer hybrid
Method: single particle / : van Eeuwen T, Taylor MS, Rout MP
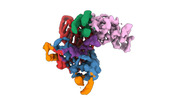
EMDB-40859: 
Structure of LINE-1 ORF2p with an oligo(A) template
Method: single particle / : van Eeuwen T, Taylor MS, Rout MP

EMDB-40305: 
Cryo-EM structure of insulin amyloid-like fibril that is composed of two antiparallel protofilaments
Method: single particle / : Wang LW, Hall C, Uchikawa E, Chen DL, Choi E, Zhang XW, Bai XC

EMDB-14619: 
Cryo-EM structure of POLRMT mutant.
Method: single particle / : Das H, Hallberg BM

PDB-7zc4: 
Cryo-EM structure of POLRMT mutant.
Method: single particle / : Das H, Hallberg BM

EMDB-25699: 
VFLIP Spike Trimer with GAR03
Method: single particle / : Sobti M, Stewart AG

EMDB-25700: 
VFLIP Spike Trimer with GAR05 FAB
Method: single particle / : Sobti M, Stewart AG, Rouet R, Langley DB

EMDB-13736: 
Cryo-EM structure of POLRMT in free form.
Method: single particle / : Das H, Hallberg BM

EMDB-13796: 
2021-10-18
Method: single particle / : Das H, Hallberg BM

PDB-7pzr: 
Cryo-EM structure of POLRMT in free form.
Method: single particle / : Das H, Hallberg BM

EMDB-13735: 
Mitochondrial DNA dependent RNA polymerase homodimer.
Method: single particle / : Das H, Hallberg BM

PDB-7pzp: 
Mitochondrial DNA dependent RNA polymerase homodimer.
Method: single particle / : Das H, Hallberg BM

EMDB-26126: 
Structure of the L. blandensis dGTPase in the apo form
Method: single particle / : Klemm BP, Sikkema AP

EMDB-26127: 
Structure of the L. blandensis dGTPase bound to dATP
Method: single particle / : Klemm BP, Sikkema AP

EMDB-26128: 
Structure of the L. blandensis dGTPase H125A mutant bound to dGTP
Method: single particle / : Klemm BP, Sikkema AP

EMDB-26129: 
Structure of the L. blandensis dGTPase H125A mutant bound to dGTP and dATP
Method: single particle / : Klemm BP, Sikkema AP

PDB-7tu5: 
Structure of the L. blandensis dGTPase in the apo form
Method: single particle / : Klemm BP, Sikkema AP, Hsu AL, Borgnia MJ, Schaaper RM

PDB-7tu6: 
Structure of the L. blandensis dGTPase bound to dATP
Method: single particle / : Klemm BP, Sikkema AP, Hsu AL, Borgnia MJ, Schaaper RM

PDB-7tu7: 
Structure of the L. blandensis dGTPase H125A mutant bound to dGTP
Method: single particle / : Klemm BP, Sikkema AP, Hsu AL, Borgnia MJ, Schaaper RM

PDB-7tu8: 
Structure of the L. blandensis dGTPase H125A mutant bound to dGTP and dATP
Method: single particle / : Klemm BP, Sikkema AP, Hsu AL, Borgnia MJ, Schaaper RM
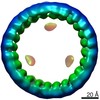
EMDB-24425: 
Circular tandem repeat protein with novel repeat topology and enhanced subunit contact surfaces
Method: single particle / : Shen BW, Stoddard BL

PDB-7rdr: 
Circular tandem repeat protein with novel repeat topology and enhanced subunit contact surfaces
Method: single particle / : Shen BW, Stoddard BL

EMDB-11978: 
Nanobody E bound to Spike-RBD in a localized reconstruction
Method: single particle / : Hallberg BM, Das H

EMDB-11981: 
SARS-CoV-spike bound to two neutralising nanobodies
Method: single particle / : Hallberg BM, Das H

PDB-7b14: 
Nanobody E bound to Spike-RBD in a localized reconstruction
Method: single particle / : Hallberg BM, Das H

PDB-7b18: 
SARS-CoV-spike bound to two neutralising nanobodies
Method: single particle / : Hallberg BM, Das H

EMDB-23118: 
Orexin Receptor 2 (OX2R) in Complex with G Protein and Natural Peptide-Agonist Orexin B (OxB)
Method: single particle / : Hong C, Byrne NJ, Zamlynny B, Tummala S, Xiao L, Shipman JM, Partridge AT, Minnick C, Breslin MJ, Rudd MT, Stachel SJ, Rada VL, Kern JC, Armacost KA, Hollingsworth SA, O'Brien JA, Hall DL, McDonald TP, Strickland C, Brooun A, Soisson SM, Hollenstein K

EMDB-23119: 
Orexin Receptor 2 (OX2R) in Complex with G Protein and Small-Molecule Agonist Compound 1
Method: single particle / : Hong C, Byrne NJ, Zamlynny B, Tummala S, Xiao L, Shipman JM, Partridge AT, Minnick C, Breslin MJ, Rudd MT, Stachel SJ, Rada VL, Kern JC, Armacost KA, Hollingsworth SA, O'Brien JA, Hall DL, McDonald TP, Strickland C, Brooun A, Soisson SM, Hollenstein K

EMDB-11980: 
SARS-CoV-spike RBD bound to two neutralising nanobodies.
Method: single particle / : Hallberg BM, Das H

PDB-7b17: 
SARS-CoV-spike RBD bound to two neutralising nanobodies.
Method: single particle / : Hallberg BM, Das H

EMDB-23018: 
SARS-CoV-2 spike in complex with nanobodies E
Method: single particle / : Hallberg BM, Das H

PDB-7ksg: 
SARS-CoV-2 spike in complex with nanobodies E
Method: single particle / : Hallberg BM, Das H

EMDB-9401: 
CH505 SOSIP.664 trimer in complex with DH522.2 Fab
Method: single particle / : Fera D, Harrison SC
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
